advancing Microscopy to Accelerate understanding of Complexity and Heterogeneity of 3D Cancer

Drug resistance is a major challenge for cancer therapy. It is believed to arise from heterogeneous cellular behaviours within tumours, where initially identical cells can mutate and adapt to diverse microenvironments. While a cancer therapy will kill most cancer cells, some “outlier” cells may evolve to “escape” the therapy and ultimately lead to recurrence of the cancer. This complex behaviour is rarely addressed in standard anti-cancer drug discovery assays that typically measure the average response of cell populations to drug candidates. Moreover, such assays usually employ cells in highly artificial contexts, such as thin 2D homogenous cell layers that are ideal for microscopy but which fail to account for the complexity that outlier cells may exploit to drive drug resistance. There is increasing interest in using more complex 3D cancer tissue models for drug discovery, such as patient-derived organoids (PDO) that better recapitulate the complexity of the in vivo context compared to conventional high throughput assays of homogenous 2D cell cultures. Unfortunately, increasing the physiological complexity of 3D cancer models changes their optical properties – making them more opaque and therefore harder to image – limiting opportunities for high throughput assays.
This CRUK Accelerator project aims to explore the trade-off between complexity of 3D cancer models and power of assays (in terms of single cell resolution and throughput) by developing modular, open source, automated instrumentation for rapid 3D imaging of such complex 3D cancer models and testing these instruments on a range of 3D cell cultures with different biological and optical properties. The emphasis on open source instrumentation is intended to enable other laboratories to replicate the capabilities that we develop in this Accelerator. We will also explore and optimise cell culture and sample preparation (e.g. labelling, mounting, clearing) techniques and share our know-how to enable researchers to optimise 3D assays to address their specific cancer biology questions.
The proposed automated 3D imaging instrumentation aims to provide quantitative single cell-resolved readouts of fluorescence biosensors reporting responses to (anti-cancer) drugs in fixed and live cell 3D cancer models arrayed in multiwell plates. Open source instrumentation will provide quasi-confocal microscopy using spinning disc scanners , oblique plane microscopy (OPM) providing light-sheet optical sectioning for fast volumetric imaging and multibeam, multiphoton microscopy for imaging deeper into 3D cell cultures recapitulating biological tissue. This new modular imaging platform will extend the commercial state-of-the-art with rapid automated 3D imaging, e.g. of cell morphology, dynamics and migration, of complex samples arrayed in multiwell plates and will provide the ability to tune performance between speed and imaging depth for larger and/or more scattering samples. We will complement open hardware configurations with software for 3D image acquisition and analysis and will develop and validate robust and reproducible protocols and SOPs for culturing and assaying 3D cell cultures including patient-derived organoids.
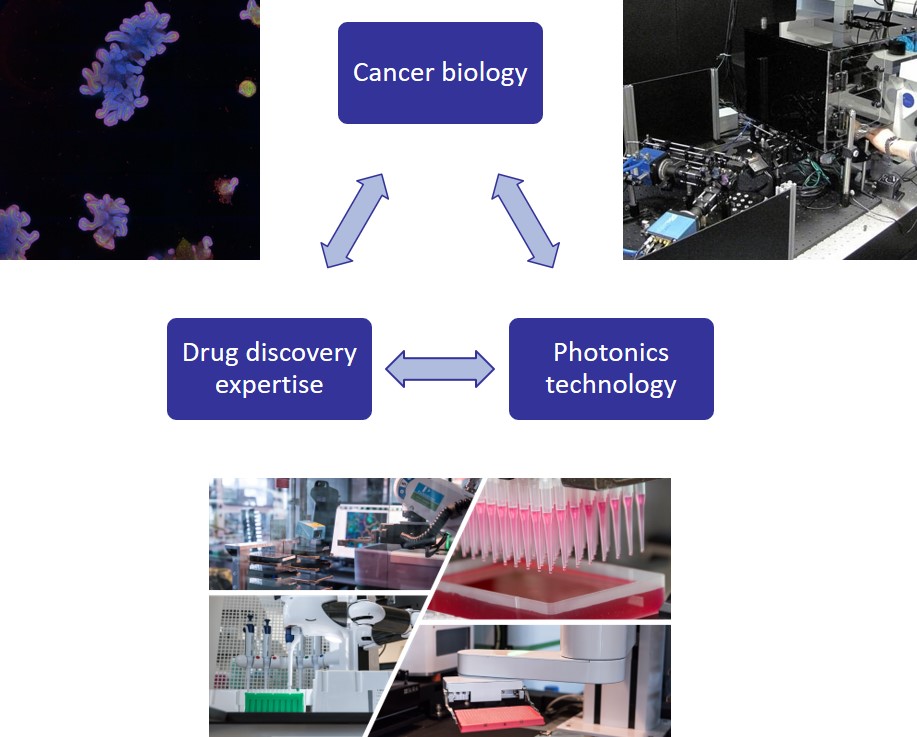
This Accelerator project combines world class cancer biology with novel photonics technologies and cutting edge high throughput imaging capabilities for drug discover
Following initial instrument development at Imperial and the Crick, we will deploy and validate the Accelerator capabilities at CRUK Edinburgh, the ICR and the IRB Barcelona through application to exemplar cancer biology assays, e.g. determining which cells within a heterogeneous cancer population are effectively killed by chemotherapy and which of the persisting cells are responsible for disease recurrence. We will also explore the role of the tumour microenvironment for quiescent resistant tumour cell sub-populations. To better understand side-effects of chemotherapy, we will assay the effects of chemotherapy on the regenerative capacity of stem cell models of normal tissue. A PhD cohort trained in these technologies will help introduce them to the cancer biology and drug discovery communities.
If you have any queries, contact us at info@mach3cancer.org
